A Home for Microbes to Share
What lives inside and on our body is the subject of emerging microbiome research. Jan Kehrmann’s team has discovered incredible things.
Von Jennifer Meina
They colonize your skin, the mucous membranes of the intestine and the lungs – milling around and multiplying everywhere: about 39 billion bacteria. Not a very pleasant thought? On the contrary, they’re essential to life. Our minuscule lodgers – bacteria, viruses, fungi, parasites, which all together make up the microbiome – have a massive influence on health and sickness.
So what do the 1,000 different species of bacteria get up to on, in and with us? What combination keeps us healthy, what makes us sick? And can the microbiome be modified so as to prevent or cure diseases? Assistant Professor Dr Jan Kehrmann from the Institute of Medical Microbiology investigates these issues. His focus: the genome of our microbe residents. ‘For roughly 15 years, the use of Next-Generation-Sequencing-technology has made it possible to have an entirely new view of microbial communities. We are inhabited by far more different types of bacteria than was previously thought. We are still only at the beginning of understanding the complex connections. But one thing is becoming increasingly clear: how important microorganisms are for us to remain healthy,’ says the senior physician.
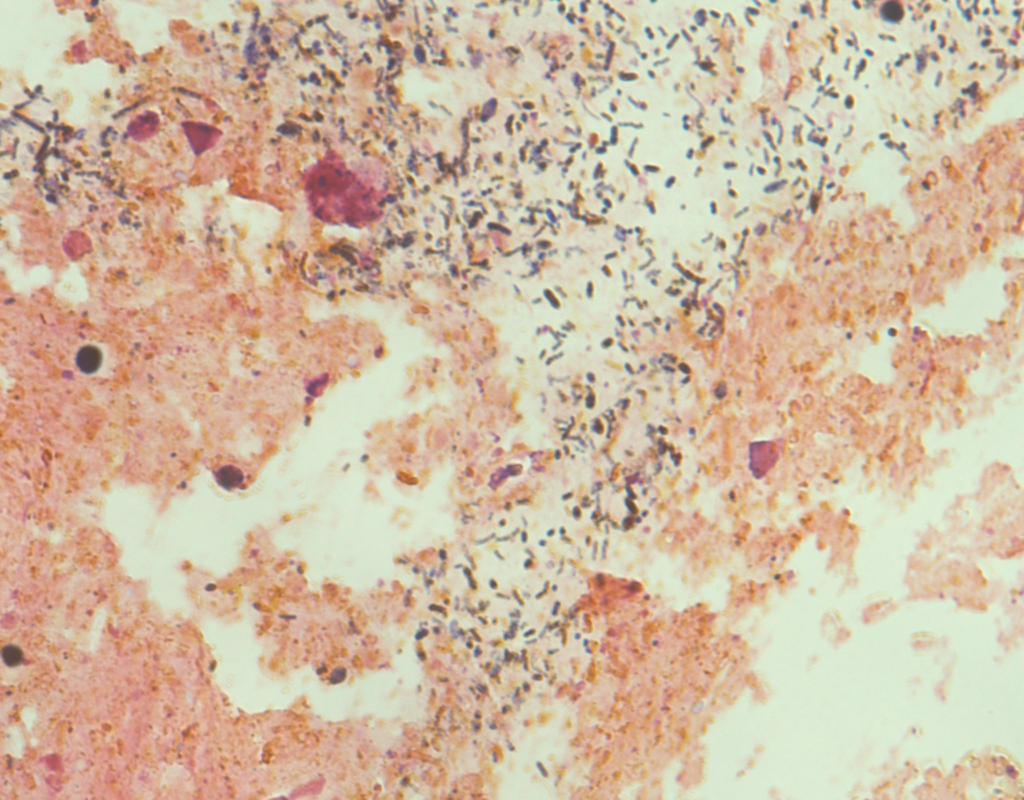
The research team is particularly interested in the intestine, where more than 99 percent of our microorganisms live. Here, bacteria not only transform the nutrients in food to make them usable by humans, but are also involved in the maturation of immune cells. They even ‘communicate’ with remote organs by means of metabolic products. In animal models, it has already been possible to reveal the connection between psychological disorders such as depression and an altered gut microbiome. Overweight and autoimmune diseases such as diabetes, multiple sclerosis or allergies have also been triggered in animal models by adjusting the gut microbiome.
»The poor image of bacteria is unjust.«
TREATMENTS OF THE FUTURE
Kehrmann emphasizes that the poor image of bacteria is unjust. Some may cause infectious diseases, but many of the various bacteria that colonize us keep us healthy. The more diverse the microbiome is, the better it seems to function. But how come? Finding the answer to this is the primary goal of the research. A recent study* by Kehrmann and his colleagues has shown that anyone who is sick with SARS-CoV-2 has a less diverse gut flora – bacteria that are associated with various inflammatory diseases proliferate, while at the same time there are fewer anti-inflammatory bacteria found in the gut flora in severe cases of Covid-19.
What does this mean? ‘It appears that healthy people have a vastly different gut microbiome than sick people. In severe cases of Covid-19 we observe an excessive immune response with increased proinflammatory cytokines and alteration of the immune cells in the blood.’ The extent to which this is caused by the human gut microbiome is not yet clear. Modeled in mice, however, it has already been possible to prove a connection for other viral lung diseases, e.g. influenza, between the gut microbiome and the immune response in the lungs, and the severity of the course of the disease. This is known as the ‘gut-lung axis’.
In addition, the researchers conjecture that the effectiveness of certain therapies varies depending on the gut microbiome. This could in future lead to analysis of gut inhabitants – the hunt for specific biomarkers – becoming as standard an examination as blood tests. ‘Doctors could see from the results who is likely to suffer a severe disease or who responds less well to specific treatments.’
UNDERSTANDING THE MECHANISMUS
The treatment response question is also the subject of the research that Kehrmann’s team is currently conducting with dermatologists from University Hospital Essen. ‘We’ve discovered that the skin microbiome changes during treatment of actinic keratosis – a precursor to non-melanoma skin cancer – and the response to treatment also varies in relation to the composition of the skin microbiome.’ Treatment is less effective in patients with a low relative abundance of corynebacteria in their microbiome, and so the disease progresses. For the future, it is important to discover whether the bacteria cause this – and what skin microbiome mechanisms influence cell growth. Depending on the result, this might lead to making specific alterations to the skin microbiome or implanting missing bacterial metabolic products before or during treatment, in order to improve the response. However, that is all still up in the air.
‘Treatment of diseases by restoring a healthy microbiome is still the exception, although diarrhoea from Clostridioides difficile, which occurs after courses of antibiotics, has been successfully treated for several years now with fecal microbiota transplantation.’ As a microbiologist Kehrmann is confident: the answers to many medical questions will probably be found in the minuscule world of the microbiome. At the same time, he warns against getting hopes up too high, ‘There are massive expectations of research into the microbiome and hundreds of studies in which fecal microbiota are transplanted in order to improve the course of the disease. They won’t all fulfil expectations, and many will be disappointing.’ Changes to the microbiome certainly cannot heal every disease. After all, a human isn’t a mouse – results from animal studies can’t be replicated one-to-one in humans.
So an important next step is to understand the mechanisms, how the microbiome influences disease or recovery – and how the disease influences the microbiome in turn.
* The study involved 212 patients, including 117 infected with SARS-CoV-2 and 95 who tested negative, that had contacted the University Medical Centre Essen on account of other ailments.

MILESTONES IN MICROBIOME RESEARCH:
Since 2005, Next-Generation-Sequencing-technology has enabled rapid analysis of the human gut microbiome. First, the bacteria must be cultivated – this used to be barely possible given some species’ difficult habitat requirements. Today, bacteria are specified quickly and easily using DNA sequencing. This technical progress has enabled scientists not only to discover the variety of billions of organisms, but also to research how closely they are linked to our health.
Main image: © Jan Kehrmann